Abstract
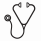
Background. Minimal residual disease (MRD) is the strongest prognostic factor in both children and adults with acute lymphoblastic leukemia (ALL). Currently, it is most widely monitored by molecular methods based on real-time-quantitative-PCR (RQ-PCR). Digital-droplet-PCR (ddPCR) and next-generation-sequencing (NGS) represent advanced tools that have the potential to overcome some limitations of standard approaches and potentially provide additional benefits. We analyzed adult ALL follow-up (FU) samples by RQ-PCR, ddPCR and NGS in order to better define the discriminating power of these novel methods.
Patients and Methods. Thirty adult ALL patients enrolled in the GIMEMA LAL 1913 protocol and their 83 FU bone marrow (BM) samples were studied. All patients received homogeneous induction/early consolidation chemotherapy, with concurrent MRD analysis at four time-points, to optimize risk classification and support risk/MRD-oriented therapy. RQ-PCR analyses followed the EuroMRD Consortium guidelines (van der Velden, 2007), ddPCR was performed as published (Della Starza, 2016; Cavalli, 2017) and NGS, as previously described (Faham, 2012; Kotrova M, 2017).
Results. By MRD RQ-PCR analysis, 19/83 samples were positive and quantifiable (Q), 9/83 were positive not-quantifiable (PNQ) and 55/83 were negative (NEG). By MRD ddPCR analysis, 27/83 samples were Q, 1/83 sample was PNQ and 55/83 proved NEG. Comparing the results of the two methods, we observed that MRD detection was concordantly positive or negative in 81% (67/83) of FU samples, while 19% (16/83) samples were classified as discordant. Most of the discordances occurred in samples with low levels of disease, i.e. PNQ or NEG: 9/83 were RQ-PCR PNQ, 4 of which were Q by ddPCR and 5 were ddPCR NEG. In the remaining 7 discordant FU samples, 5 were RQ-PCR NEG/ddPCR Q, 1 sample was RQ-PCR Q /ddPCR NEG and 1 sample was RQ-PCR NEG/ddPCR PNQ. The use of ddPCR significantly reduced the proportion of PNQ samples if compared to RQ-PCR - 1/83 (3%) vs 9/83 (15%) - respectively (p=0.0179), increasing the proportion of Q samples: 27/83 (33%) vs 19/83 (23%). It is worth noting that ddPCR also quantified the levels of disease in 9% (5/55) of samples, that were RQ-PCR NEG (Table 1).
MRD analysis was also performed by NGS in 41 samples from 15 patients: 18/41 samples proved Q and 23/41 were NEG. Comparing the MRD detection obtained by both ddPCR and NGS, we observed a concordant result in 98% (40/41) of samples; only 1 sample was ddPCR NEG and NGS Q with a MRD level of 1x10-5. The concordance between RQ-PCR and NGS was 78% (32/41 samples). Moreover, among these 41 samples 9 (from 7 patients) were discordant between RQ-PCR and ddPCR in the first comparative analysis: in 4 RQ-PCR-NEG FU samples, 3 were Q by both ddPCR and NGS, 1 was ddPCR NEG and NGS Q, with a MRD level of 10- 5; 1 subsequent relapse was observed; 4 FU samples that were RQ-PCR-PNQ/ddPCR-Q, were Q also by NGS; 1 subsequent relapse was observed. Finally, 1 RQ-PCR-PNQ sample was negative by both ddPCR and NGS, and no recurrence has so far been observed. Moreover, in the cohort of samples analyzed only by RQ-PCR and ddPCR, in 1 RQ-PCR NEG/ddPCR Q sample a relapse was observed, while the only case that was RQ-PCR Q/ddPCR NEG has so far not relapsed. Notably, 2 of the 3 relapses were documented in patients who were, at decisional treatment TPs, RQ-PCR PNQ or NEG and ddPCR/NGS Q.
Conclusions. When MRD levels are very low, it can be difficult to dissect if the not-quantifiable signal observed by PCR is due to few residual leukemic cells or to a non-specific amplification of normal DNA. The superior sensitivity and accuracy of ddPCR and NGS could be instrumental to univocally define these samples, which presently represent a problematic gray area in the clinical practice of MRD-driven protocols and might be associated with clinical relapse: indeed, among 83 FU samples analyzed we observed 3 relapses, whose FU samples were classified as PNQ or NEG by RQ-PCR, but proved Q by ddPCR and/or NGS. At variance, no relapses were recorded in patients whose FU samples were defined RQ-PCR-PNQ, but proved ddPCR/NGS NEG. Moreover, in 2/3 relapsed cases the change of MRD status (PNQ or NEG vs Q) could have led to a switch in risk classification and therefore in a treatment change. Further studies with a larger number of discrepant cases and a longer FU time will allow to conclusively define the clinical application and implication of these new methods.
Chiaretti:Shire: Consultancy; Pfuzer: Consultancy; Amgen: Consultancy; Incyte: Consultancy. Foà:NOVARTIS: Speakers Bureau; ROCHE: Other: ADVISORY BOARD, Speakers Bureau; CELTRION: Other: ADVISORY BOARD; ABBVIE: Other: ADVISORY BOARD, Speakers Bureau; CELGENE: Other: ADVISORY BOARD, Speakers Bureau; JANSSEN: Other: ADVISORY BOARD, Speakers Bureau; INCYTE: Other: ADVISORY BOARD; AMGEN: Other: ADVISORY BOARD; GILEAD: Speakers Bureau.
Author notes
Asterisk with author names denotes non-ASH members.
This icon denotes a clinically relevant abstract

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal